This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
protein interaction networks
Proteins, derived from RNA, interact with various other proteins in maps known as interaction networks. Through identification of these protein networks, cellular processes can be better understood in addition to a more thorough understanding of what proteins may be involved in certain genetic diseases [1]. Though various experiments can be performed to better understand these interactions, one specific test is using a Tandem Affinity Purification, or TAP, to bind bait proteins, alter them--for example, with CRISPR--and observe the effects on other proteins [2]. Another option is to utilize a Yeast-2-Hybrid screen. In turn, upregulation or downregulation of these bait proteins should interfere with other wild type protein functions, thus creating the interaction network.
Once these networks are established, they are compiled into STRING, a database containing interaction networks for thousands of proteins. After searching with a protein name, STRING then suggests various organisms and protein homologs for further specification. After choosing an organism, the interaction network is shown.
Once these networks are established, they are compiled into STRING, a database containing interaction networks for thousands of proteins. After searching with a protein name, STRING then suggests various organisms and protein homologs for further specification. After choosing an organism, the interaction network is shown.
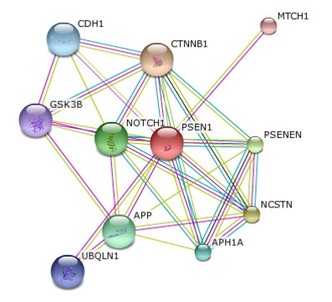
PSEN1 interaction network
|
Using the aforementioned STRING database, the interaction network for PSEN1 was identified. After sorting the proteins by gene ontology, PSEN1 was highlighted as being influential in cell adhesion, cell signalling, cellular transport and the gamma-secretase sub-unit. Despite this, one protein did not directly correspond with any other in the network, specifically UBQLN1.
Known as Ubiquitin-1, STRING correlated UBQLN1 with PSEN1 accumulation in humans. As a buildup in mutated PSEN1 would lead to the known phenotypic symptoms of Alzheimer's, further research was conducted on this knew protein. |
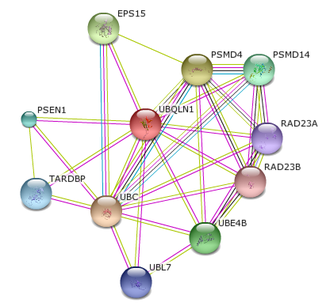
UBQLn1 interaction network
|
Known to impact PSEN1 accumulation, STRING was once again consulted to identify the interaction network for UBQLN1. Though still corresponding with PSEN1, UBQLN1 was found to interact with other proteins in regards to cellular processes such as DNA binding, the proteosome, growth factors and ubiquitination.
This dual interaction network not only further derives the complexity of Alzheimer's disease (through identifying new cellular functions that may be altered in the disease state), but also reassures what modern Alzheimer's research should focus on. |
discussion
Mentioned above, the protein interaction networks were crucial to identifying novel research focuses for future Alzheimer's experiments. Though the initial network for PSEN1 may be smaller than other protein interaction networks, the earlier results highlight just how complex Presenilin 1 and Alzheimer's disease really are. These identified interaction networks will prove essential to understanding additional cellular functions affected through Alzheimer's. Lastly, these results indicate that future research should pertain to UBQLN1, as the role in PSEN1 accumulation could be directly related to the development of Alzheimer's.
REFERENCES