This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
post-translational modifications
Post-translational modifications, as expected, occur after translation. These modifications can affect proteins in a variety of ways: targeting for degradation, altering interactions, gene expression effects, differentiation and other various cellular processes. Of the many post-translational modifications that a protein can undergo, the most common and well-understood is phosphorylation [1].
NetPhos 2.0 is a website used to predict phosphorylation sites of a protein based on specific amino acids. As such, there are three main amino acids that NetPhos 2.0 looks for: threonine, tyrosine and serine. After entering the protein FASTA sequence, the database gives each amino acid a score based on 'phosphorylation potential'--a value between zero and one. In this case, the closer of a score to 'one', the more likely that amino acid is phosphorylated.
NetPhos 2.0 is a website used to predict phosphorylation sites of a protein based on specific amino acids. As such, there are three main amino acids that NetPhos 2.0 looks for: threonine, tyrosine and serine. After entering the protein FASTA sequence, the database gives each amino acid a score based on 'phosphorylation potential'--a value between zero and one. In this case, the closer of a score to 'one', the more likely that amino acid is phosphorylated.
netphos 2.0 results
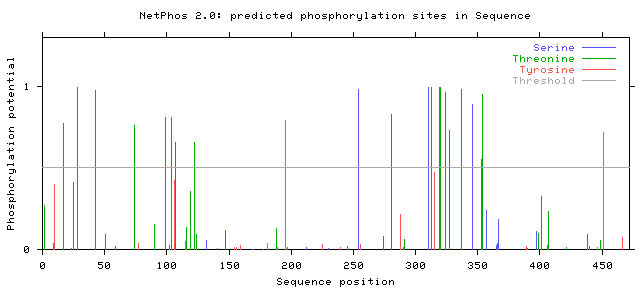
Using NetPhos 2.0 to obtain plausible phosphorylation sites, human PSEN1 was predicted to have 8 threonine, 2 tyrosine and 11 serine phosphorylation sites. From here, the NetPhos 2.0 results can be compared to the protein domain of PSEN1 to determine which phosphorylation sites are most pertinent to the active domain. For additional information regarding the PSEN1 protein domain, click here.
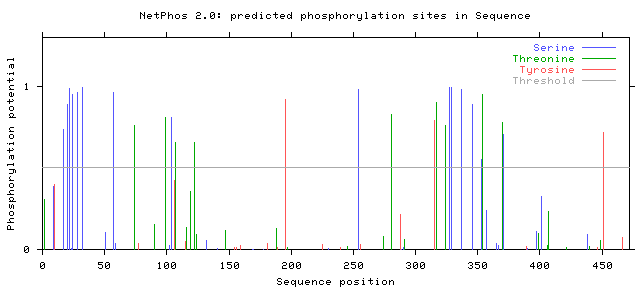
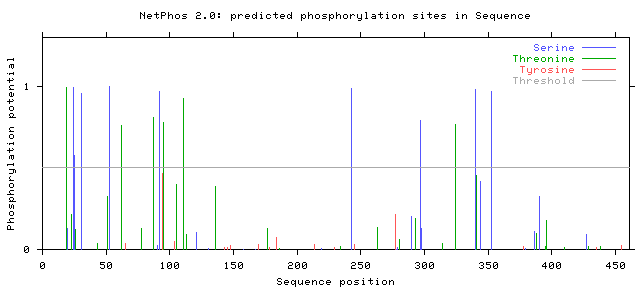
In addition, the results from human PSEN1 can be compared to NetPhos 2.0 results for the chosen model organisms, mice and zebrafish. The results are seen below. Click on each for a larger picture:
In addition, the results from human PSEN1 can be compared to NetPhos 2.0 results for the chosen model organisms, mice and zebrafish. The results are seen below. Click on each for a larger picture:
Protein domain map for human PSEN1.
NetPhos 2.0 results for humans.
discussion
Through the various NetPhos 2.0 results included above, pertinent phosphorylation sites for human PSEN1 can be identified. With the comparison to the included protein domain, certain amino acid sites may prove to be more essential to wild type PSEN1 function than others. One example of this is found at serine-320, indicated by the red line on the aforementioned protein domain included above. Because this site appears to be the most conserved between humans, mice and zebrafish, this phosphorylation may be essential to regular PSEN1 function and regulation of Alzheimer's disease. This is further established by the location of serine-320 in the active presenilin domain. For more information on this development, please view the 'Future Directions and Conclusions' page.
REFERENCES