This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
Protein motifs
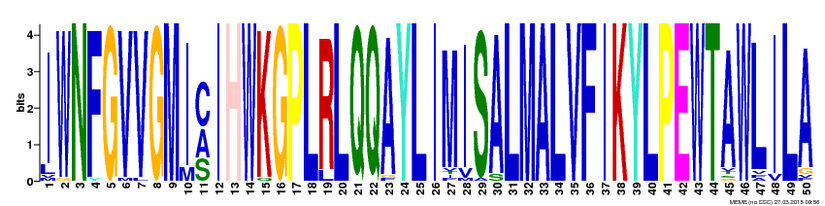
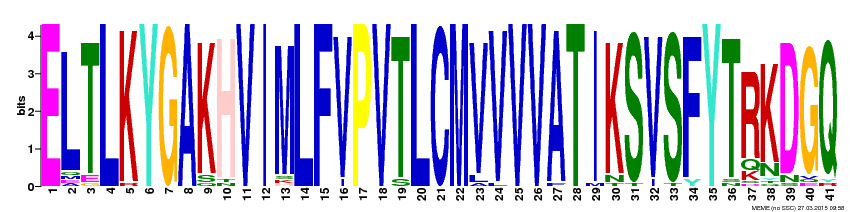
Protein motifs are used to estimate and predict protein function through attempts to predict the products of each gene [1]. As several motifs can appear throughout a sequence, more than one motif can generally be produced. To account for such, there are several different databases that can be used to produce protein motifs, such as MEME. Such motifs are then useful to identify how conserved the gene sequences are across organisms [2]. The larger the letter, the more conserved that region is across various species.
Across all the species studied in the proposed phylogenetic trees, three possible protein motifs are listed below. While none are 100% accurate, they provide an indication for commonality across 41 different organisms. See the motifs below:
Across all the species studied in the proposed phylogenetic trees, three possible protein motifs are listed below. While none are 100% accurate, they provide an indication for commonality across 41 different organisms. See the motifs below:
protein domains
Protein domains are used in genetics research to illustrate the location of functional units of a protein. Each functional unit/domain is used for a specific function or role inside the cell [3]. Especially useful in relatively conserved proteins, domains allow for a direct comparison of how a protein appears in one organism over another. In the case of presenilin 1, the percentage of how identical each domain is to the human variant of the protein can be established. To obtain these domains, various databases can be referred to, such as SMART, Prosite, or Pfam, which can gather and sort protein domains based on protein sequences entered.
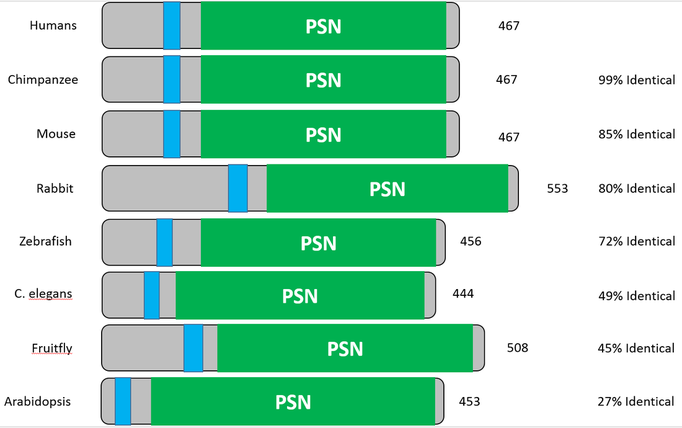
Seen below are the protein domains established using SMART. After entering the protein sequence into the database, the domain architecture was created. In order to understand the domains, they have been color coordinated. The green, PSN region of the domain indicates where presenilin is found within the PSEN1 protein. The light blue, smaller region depicts a transmembrane domain--an intrinsic property of the protein.
Other helpful information includes the name of the organism associated with each protein domain, found to the left, the amount of amino acids found in the protein, found to the right, and the percentage of how identical each domain is to the human homolog of the protein. See the results below:
Seen below are the protein domains established using SMART. After entering the protein sequence into the database, the domain architecture was created. In order to understand the domains, they have been color coordinated. The green, PSN region of the domain indicates where presenilin is found within the PSEN1 protein. The light blue, smaller region depicts a transmembrane domain--an intrinsic property of the protein.
Other helpful information includes the name of the organism associated with each protein domain, found to the left, the amount of amino acids found in the protein, found to the right, and the percentage of how identical each domain is to the human homolog of the protein. See the results below:
Discussion
The above data illustrates just how conserved PSEN1 is throughout various organisms and species. Seen through both the motifs and domains, PSEN1 remains almost identical through most mammals--an unshocking observation knowing how presenilin is actively involved in neuronal function and regulation. Each organism above only has the PSN domain and a transmembrane domain, backing the claim of how rigid the structure of the presenilin 1 protein is. However, this information is also very insightful, as it reassures which organisms should be used to model Alzheimer's disease research, namely both mouse and zebrafish.
REFERENCES
[1] http://www.proteinstructures.com/Structure/Structure/protein-motifs.html
[2] http://www.barabasilab.com/pubs/CCNR-ALB_Publications/200310-00_NatureGen-Evolutionary/200310-00_NatureGen-Evolutionary.pdf
[3] http://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains
[2] http://www.barabasilab.com/pubs/CCNR-ALB_Publications/200310-00_NatureGen-Evolutionary/200310-00_NatureGen-Evolutionary.pdf
[3] http://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains