This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
Chemical genetics
Chemical genetics are increasingly useful in establishing how certain drugs or treatments affect genes and proteins. Using databases such as PubChem and Chembank, various chemicals that should react with the gene of interest should be identified--done by testing a chemical's binding properties as well as the observed phenotypic result [1]. As these databases are relatively large, finding a small molecule, or chemical treatment to certain genes should be possible, as is the case with PSEN1 and Alzheimer's. Whilst these databases only contain known chemical treatments, there is always the possibility of currently unknown treatments. This makes studying chemical genetics increasingly important, as by finding compounds that can target and treat certain mutated genes, is an essential component to dissipating disease symptoms, like the memory-associated ones of Alzheimer's.
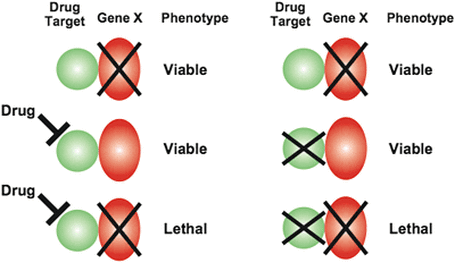
Seen below is a typical chemical genetic experiment. Illustrated is how different drug targets could bind to a gene, causing varying phenotypic results dependent on if the drug treatment was successful or not. Continue below:
Seen below is a typical chemical genetic experiment. Illustrated is how different drug targets could bind to a gene, causing varying phenotypic results dependent on if the drug treatment was successful or not. Continue below:
Figure 1. A typical chemical genetic experiment, illustrating the success of each respective test.
PSEN1 chemical treatments
When searching for PSEN1 as a target in the PubChem database, 148 different compounds were identified for human presenilin 1 [2]. While indicating that these chemicals can bind to human PSEN1, none have been successful in preventing upregulated neurotoxicity under mutated PSEN1 or preventing amyloid plaque and neurofibrillary tangle formation. As such, these chemicals could be tested in our mice and zebrafish model organisms to test the effects on similarly conserved presenilin 1 regions.
In conjunction with the protein interaction networks, several chemical genetic could be performed on the associated proteins of PSEN1--observing if any of those may have a downregulatory effect on APP production in organisms with mutated PSEN1. While a long shot, the relatively rapid capacity of both mice and zebrafish should allow this chemical genetic screen to be performed.
In conjunction with the protein interaction networks, several chemical genetic could be performed on the associated proteins of PSEN1--observing if any of those may have a downregulatory effect on APP production in organisms with mutated PSEN1. While a long shot, the relatively rapid capacity of both mice and zebrafish should allow this chemical genetic screen to be performed.
|
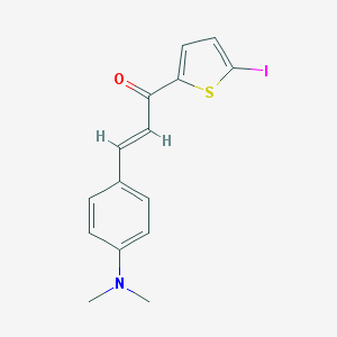
Figure 2. Compound CID 17757413 used in model mice Alzheimer's research.
|
In addition, one chemical component found in PubChem while performing a bioassay for Alzheimer's was compound CID 17757413. This chemical compound, utilized in May of 2014, proved to reduce the binding APP in mice with mutated PSEN1 [3]. Doing so allowed for the suppression of Alzheimer's disease symptoms and the support of healthy brain function.
As this compound has only been tested for relative efficiency in mice, it is currently known what effects this compound would have on humans. I would suggest to test this chemical on PSEN1 mutated zebrafish--if the results appear promising for Alzheimer's treatment, it could eventually proceed to human trials. Being able to test chemical compounds in various organisms, there is an underlying sense of hope regarding Alzheimer's and other genetic diseases. Rapid testing of compounds like CID 17757413 only increase the understanding and capacity for disease prevention. Research is the first step. |
REFERENCES
Figure 1. http://sites.utoronto.ca/boonelab/research_projects/chemical_genetics/interaction_networks/index.shtml
Figure 2. http://pubchem.ncbi.nlm.nih.gov/compound/17757413?from=summary
[1] http://www.pnlab.org/publications/documents/5700853a.pdf
[2] http://pubchem.ncbi.nlm.nih.gov/targets/?id=1709856
[3] http://pubchem.ncbi.nlm.nih.gov/assay/assay.cgi?aid=300749
Figure 2. http://pubchem.ncbi.nlm.nih.gov/compound/17757413?from=summary
[1] http://www.pnlab.org/publications/documents/5700853a.pdf
[2] http://pubchem.ncbi.nlm.nih.gov/targets/?id=1709856
[3] http://pubchem.ncbi.nlm.nih.gov/assay/assay.cgi?aid=300749